LinkML for Scientific Software: Schema Workflows with Real Leverage
LinkML for Scientific Software: Schema Workflows with Real Leverage
These blog posts are intended to provide software tips, concepts, and tools geared towards helping you achieve your goals. Views expressed in the content belong to the content creators and not the organization, its affiliates, or employees. If you have any questions or suggestions for blog posts, please don’t hesitate to reach out!
Introduction
Scientific software pipelines often break because of interfaces instead of algorithms. LinkML lets you define a single schema once and then generate validators, Python models, and visualizations from that same source. This gives you a typed contract for domain metadata, sample sheets, and analysis outputs, with less drift across scripts, notebooks, and downstream tools. It also gives you a solid mental model to communicate and understand scientific problem spaces.
Quick Terms
-
Data model: the conceptual shape of your domain objects and relationships (for example, a queue has many orders). -
Specification: the written rules for how data should look and behave. -
Schema: the machine-readable version of specifications used for validation and tooling. -
Ontology: a shared vocabulary of meaning that connects models, specifications, or schemas to broader community concepts.
Why Use Schemas Instead of Ad-hoc Code or Types?
Without a schema, pipelines often rely on implied conventions:
- IDs are “supposed” to follow some pattern.
-
status(as an example of a field) is “supposed” to be one of a few values. - Numeric fields are “should” to have sensible bounds.
Those assumptions are typically encoded in multiple places and diverge over time. LinkML centralizes those constraints into one model and reuses it across tooling.
A Minimal Viable Research Software Beverage (MVRSB) Schema in LinkML
# This is the schema's unique name on the web.
id: https://example.org/coffee-queue
# This is the short local name for the schema.
name: coffee_queue
# Shortcuts for longer URLs.
prefixes:
ex: https://example.org/
linkml: https://w3id.org/linkml/
# Pull in built-in scalar types such as string/integer.
imports:
- linkml:types
# Use `ex` if no prefix is given.
default_prefix: ex
# Use text/string if no type is given.
default_range: string
# Main object types in this schema.
classes:
# The top object in each data file.
CafeQueue:
tree_root: true
# Fields on CafeQueue.
attributes:
# Unique ID for the queue.
queue_id:
identifier: true
# A list of order objects.
orders:
# Each item in `orders` must be an Order.
range: Order
# This field is a list.
multivalued: true
# Keep the list directly inside CafeQueue.
inlined_as_list: true
# The object type for each coffee order.
Order:
attributes:
# Unique ID for each order.
order_id:
identifier: true
drink:
required: true
# Status must be one of the allowed status words below.
status:
range: OrderStatusEnum
required: true
# Allowed word lists.
enums:
OrderStatusEnum:
permissible_values:
queued:
brewing:
ready:
In a few lines, we built a tiny but real data schema: each queue has orders, each order has required fields, and status values come from a controlled list.
LinkML Building Blocks
- Start with a
schema: this is the full document that holds everything below. - Inside the schema, define
classes: these are your object types, likeCafeQueueandOrder. - Inside each class, define
slots(here written asattributes): these are the fields on that object, likeorder_id,drink, andstatus. - Use
rangeon a slot to say what kind of value it accepts:-
range: stringmeans plain text. -
range: OrderStatusEnummeans one value from a controlled list. -
range: Ordermeans a nested object of another class.
-
- Use
enumsto define those controlled lists, likequeued,brewing, andready. - Use slot rules like
required: trueandidentifier: trueto enforce structure and uniqueness. - Use
prefixesplus ontology mappings (class_uri,slot_uri,meaning) when you want semantic interoperability across datasets.
Class Linking
Inside one schema file, you can link to additional schema using range: OtherClassName.
For example, the following schema uses the earlier CafeQueue class.
classes:
Cafe:
tree_root: true
attributes:
cafe_id:
identifier: true
queue:
range: CafeQueue
required: true
inlined: true
CafeQueue:
attributes:
queue_id:
identifier: true
This is the simplest way to model nested objects.
Composing Multiple Schema Files
Across files, linking works the same way, and imports stitches files together.
Use one top-level entrypoint and generate from that file.
Simple layout:
-
examples/linkml/modular/common.yaml: shared prefixes and enums. -
examples/linkml/modular/order.yaml:Orderclass. -
examples/linkml/modular/queue.yaml:CafeQueueclass that referencesOrder. -
examples/linkml/modular/cafe.yaml: top-levelCafeclass that referencesCafeQueue.
Minimal import chain:
# cafe.yaml
imports: [queue]
# queue.yaml
imports: [common, order]
Generate code from the top-level file:
uv run --with linkml linkml generate pydantic \
examples/linkml/modular/cafe.yaml \
> examples/linkml/modular/coffee_cafe_pydantic.py
Use generated types normally:
from coffee_cafe_pydantic import Cafe
cafe = Cafe(**payload)
Pydantic Tie-in
LinkML can generate Pydantic models directly. Pydantic turns raw dictionaries into typed Python objects and raises clear errors when required fields are missing or values are invalid.
# generate pydatanic models from a linkml schema yaml file
uv run --with linkml linkml generate pydantic \
examples/linkml/omics_study_schema.yaml \
> examples/linkml/coffee_queue_pydantic.py
Then use generated models in your app:
from coffee_queue_pydantic import CafeQueue
# create a queue from the pydantic model
queue = CafeQueue(**payload) # raises on invalid data
That pattern keeps LinkML as the source of truth while giving you first-class Pydantic ergonomics in APIs and services.
Ontology Alignment
LinkML can keep your schema lightweight while still attaching shared meaning through ontology terms. This helps when multiple teams or datasets need to interpret fields the same way. In practice, this makes cross-dataset joins, search, and reuse in graph/RDF workflows much more reliable.
prefixes:
# ex here represents an external example prefix
ex: https://example.org/
schema: http://schema.org/
classes:
Order:
class_uri: ex:Order
attributes:
drink:
# https://schema.org/recipeIngredient
slot_uri: schema:recipeIngredient
enums:
OrderStatusEnum:
permissible_values:
queued:
meaning: ex:queued
brewing:
meaning: ex:brewing
ready:
meaning: ex:ready
-
class_uri: says “this class matches this shared concept.” -
slot_uri: says “this field means this shared property.” -
meaning: says “this enum label corresponds to this controlled term.”
Visualizing the Model
You can render the same schema as either Mermaid ER or PlantUML:
uv run --with linkml linkml generate erdiagram \
examples/linkml/omics_study_schema.yaml \
> examples/linkml/coffee_queue_schema.mmd
uv run --with linkml linkml generate plantuml \
examples/linkml/omics_study_schema.yaml \
> examples/linkml/coffee_queue_schema.puml
Generator docs:
The outputs are useful in docs and design reviews because teams can quickly verify structure before implementation details distract the conversation.
Generated Mermaid (from linkml generate erdiagram on this schema):
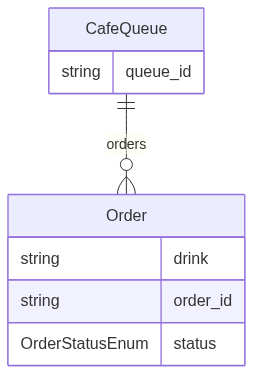
Generated PlantUML (from linkml generate plantuml on this schema):
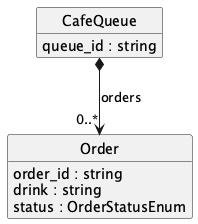
Useful for Bioinformatic Research Software Engineering
- The same pattern maps cleanly to sample sheets and assay metadata.
- Controlled vocabularies for conditions, assays, and cohorts become explicit enums.
- Provenance fields (batch, instrument, protocol version) can be required instead of implied.
- The schema stays reusable across notebooks, CLIs, and workflow engines.
- Generated artifacts make handoffs to warehouse or graph layers less brittle.
Caveats
- Schema design still takes thought; LinkML reduces drift, not modeling complexity.
- Very large legacy payloads may need an incremental migration plan.
- Teams should agree on schema versioning strategy early.
Run the Example Locally
The runnable companion demo for this post is in examples/linkml/linkml_quickstart_demo.py.
uv run examples/linkml/linkml_quickstart_demo.py
